New Genetic Variants Linked to Aggressive or Benign MS Disease Course, Study Shows
Written by |

Genetic variants in the CPXM2, IGSF9B, and NLRP9 genes were found to potentially shape the disease course of multiple sclerosis (MS), and may be used as biomarkers to identify those with an aggressive or benign type of disease, a DNA sequencing study shows.
The study, “Exome sequencing study in patients with multiple sclerosis reveals variants associated with disease course,” was published in the Journal of Neuroinflammation.
Several genetic association studies have uncovered many MS risk genes, including the well-characterized genetic variant HLA-DRB1*15:01, the strongest known genetic risk factor for MS.
However, little is known about genes that influence MS disease course. The way MS evolves varies considerably between patients, and over time. Some may experience episodes of relapse followed by periods of remission, while others experience a disease that progressively worsens. In addition, in more severe cases, the disease symptoms are markedly aggravated within a few years, while in MS cases considered benign, symptoms get worse at a slower pace.
The underlying cause of this variability is unknown, but there is scientific evidence suggesting that genetic variants may play a role.
To address this question, a team led by researchers at Hospital Universitari Vall d’Hebron, Universitat Autònoma de Barcelona in Spain conducted a DNA sequencing study to identify genes specifically associated with MS disease course.
First, they sequenced all the protein-coding genes (exome sequencing) of a group of 20 MS patients, 10 of whom had a disease course classified as aggressive and 10 characterized as having a benign disease course.
Connect with other patients and share tips on how to manage MS in our forums!
Benign MS was defined as having an Expanded Disability Status Scale (EDSS) score of three or lower over the course of 15 or more years from disease onset, and never having received MS therapies. An aggressive disease course was determined as having reached an EDSS score of six or higher within the first five years of disease onset, regardless of having received treatment.
From this first screen, researchers identified a list of 16 genetic variants — called single-nucleotide polymorphisms (SNPs) — potentially associated with either more aggressive or benign disease manifestations.
To validate the association of these genetic variants with disease course, they were assessed in two independent groups of MS patients: the first with 194 MS patients, 107 with benign and 87 with aggressive symptoms; the second with 257 patients, of whom 224 patients had benign symptoms and 33 aggressive disease courses.
From these studies, two genetic variants were validated as being associated with MS disease course, namely rs28469012 and rs10894768. The first is located in the gene CPXM2 (carboxypeptidase X, M14 family, member 2) and was associated with an aggressive disease course. The other, rs10894768, appeared in the gene IGSF9B (immunoglobulin superfamily member 9B) and was linked with a benign manifestation.
Additionally, another genetic variation called rs10423927 showed a trend, which was not statistically significant, toward association with a benign MS course. This variant is located in the gene NLRP9 (NLR family pyrin domain containing 9).
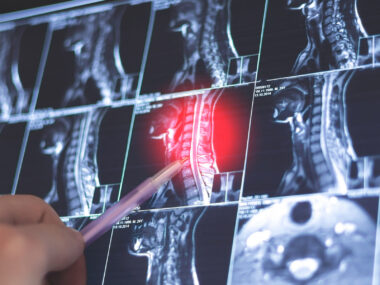
To explore whether these genetic variants could play specific roles in the formation of MS lesions at the central nervous system (CNS; spinal cord and brain), researchers investigated the expression of the proteins corresponding to the three genes in brain tissue from patients.
Specifically, they looked at the expression of IGSF9B, CPXM2, and NLRP9 proteins in chronic active lesions of brain samples from four MS patients.
IGSF9B proteins were present in astrocytes — star-shaped cells important for providing nutrients, support, and protection to the central nervous system — and macrophages/microglial cells — a type of immune cell present in the brain and spinal cord — whereas CPXM2 and NLRP9 proteins were restricted to brain macrophages/microglia.
IGSF9B and CPXM2 genes are known to play a role in the integrity of synapses, or the point of communication between nerve cells, which supports the need for additional studies “to explore at the CNS level the potential functional consequences of the reported polymorphisms associated with MS disease course,” the researchers wrote.
They concluded that “genetic variants located in CPXM2, IGSF9B, and NLRP9 have the potential to modulate disease course in MS patients.” In addition, the team believes that these genetic variants “may be used as disease activity biomarkers to identify patients with divergent disease courses.”

Vincent A Russotto
I have has MS since July 08, and never has a relapse
Royce ONan
Have your MRIs changed over the years? Are you on any treatment?
John Coccia
How about writing these in laymen term's (like book for dummies) So that Some of us don't have to spend all day trying to look up and decipher all of these medical term's. I know that it would be helpful to me.
Jim
I completely understand that. The second book I write about multiple sclerosis is going to be about using simpler terms and what it means to experience it in life.
Jana
It can be difficult to understand all of the terms used here. I have a science background and I do appreciate having the details as opposed to having told to me what they want me to know.
However, I think it would be beneficial to have a conclusion in lay terms. A suggestion is to read the beginning. Read up to where you understand and skip all the details and don’t forget the conclusion. That may be helpful.
Sally Calneggia
So true - it is so helpful for those of us trying to do the very best we can to stay well but it is very difficult when a lot of your articles only make sense and can be fully understood by those in the medical profession. I have had MS for 5 years now and have a very big belief in also following a very healthy diet, keeping fit and meditating but went onto Gilenya after being diagnosed. I have had no relapses since then. So I am 50/50 with what I believe is keeping me well but I truly believe food is also medicine and has a huge impact on how it inflames or protects our immune system. I am also so grateful for the advancements and treatments that are now available. My philosophy with my MS has been driven by reading both of George Jelenik’s books and most definitely made the most sense to me. He did explain everything so easily for those of you wanting a common sense approach.
Kyle
Is genetic testing for these alterations readily available? What are some options for either full exome or select gene sequencing?