Scientists Build Map of Toxic Immune Cells Contributing to Neurodegeneration in MS
Written by |
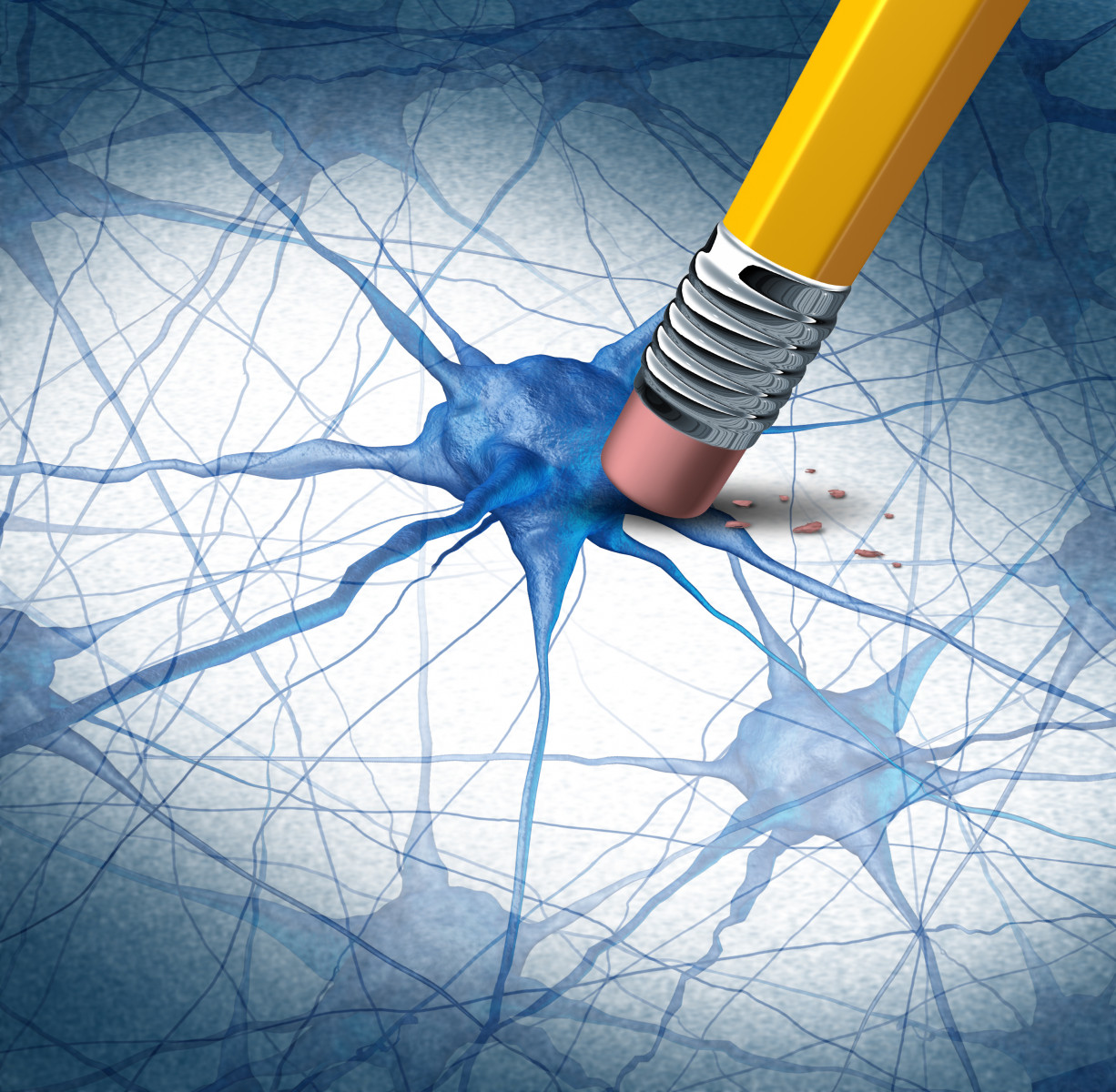

Scientists have built a map of the toxic immune cells that contribute to neurodegeneration in multiple sclerosis (MS). Their findings may open the door to the development of new medications that protect the brain from the effects brought on by these harmful immune cells.
Results were reported in the study, “Transcriptional profiling and therapeutic targeting of oxidative stress in neuroinflammation,” published in the journal Nature Immunology.
Neurodegeneration is the central hallmark of many inflammatory and neurological disorders, including MS, amyotrophic lateral sclerosis (ALS), Alzheimer’s, and Parkinson’s disease. Oxidative stress, a phenomenon in which cells are progressively damaged due to the presence of high levels of oxidant molecules, or reactive oxygen species (ROS), is one of the contributors to neurodegeneration.
In the case of MS, it has been shown that microglia — cells that support and protect neurons, and are considered the immune cells of the brain — produce large amounts of ROS that contribute to oxidative stress during the early phases of the disease, resulting in progressive brain damage. However, it was not known how immune cells controlled the production of these toxic compounds.
Now, investigators at Gladstone Institutes and their collaborators developed a technique that enables them to track when specific immune cells, which are known to produce large amounts of ROS in the central nervous system (CNS, the brain and spinal cord), activate certain genes, including those that could be involved in the production of ROS.
The new method, called Tox-seq, combines single-cell RNA-sequencing technology with a labeling technique that allows researchers to track specific types of ROS-producing immune cells, and at the same time know which genes are being “switched on” or “off” at specific time intervals.
(Note: RNA-sequencing is a technique that allows scientists to examine all RNA molecules produced from active genes in a cell or tissue.)
When applied to mice with experimental autoimmune encephalomyelitis (EAE) — a disease that mimics MS in humans — Tox-seq allowed researchers to build a map containing the genetic signature of all immune cells that contributed to the buildup of toxic ROS in the animals’ CNS.
Using this technique, the team discovered that a small sub-group of microglia, which corresponded to approximately 10% of cells, were responsible for triggering oxidative stress in the animals’ CNS, along with other sub-types of immune cells that sometimes had access to the brain.
They also found that the genetic signature of these microglia matched that of cells that previously had been thought to cause brain damage in patients with progressive forms of MS.
Moreover, Tox-seq showed that ROS-producing microglia activated genes that promoted blood clotting, which also can lead to microglia activation, brain inflammation, and oxidative stress.
“This is the first time we have evidence that coagulation and oxidative stress are at work in the same immune cells in the brain,” Katerina Akassoglou, PhD, senior investigator at Gladstone Institutes, professor of neurology at University of California San Francisco, and senior author of the study, said in a press release. “It’s a vicious cycle between the two processes.”
The team then explored how Tox-seq could identify compounds that could be used therapeutically to reduce oxidative stress in the CNS.
After screening a library of 1,907 compounds, researchers discovered 128 that were able to prevent microglia activation induced by fibrin, a blood-clotting protein. However, they still did not know which of the 128 had a specific effect on oxidative stress.
“At this point, our solution was to computationally overlay the oxidative stress gene signature identified by Tox-seq with previously published drug-target pathways,” said Jae Kyu Ryu, PhD, staff research scientist at Gladstone Institutes, and co-first author of the study. “This type of overlay had never been done before for oxidative stress pathways.”
The combination of Tox-seq data with information from previous screenings allowed researchers to zero in on compounds that could be relevant for oxidative stress.
By doing so, they found a compound called acivicin that was able to block an enzyme that destroyed glutathione, an antioxidant substance that neutralizes ROS. This meant that acivicin potentially could reduce oxidative stress by preventing glutathione from being destroyed.
The team’s hunch was confirmed. When investigators treated cells in a lab dish and EAE mice with acivicin they found that microglia activation was prevented in cells, and in animals acivicin suppressed the onset of neurological symptoms — even in those already at advanced, chronic stages of the disease.
“This was exciting because it told us that oxidative stress may be a key driver in maintaining the clinical severity of MS, not just in the initial nerve damage,” said Ryu.
Although acivicin itself may not be a promising therapeutic candidate to treat MS due to its toxic side effects, the discovery of its mechanism of action and the potential therapeutic implications of controlling oxidative stress may steer the development of new medications for the disease.
“We now have a map of the toxic immune cells in the brain that drastically changes our understanding of how disease develops and how it can be treated,” Akassoglou said. “The map of toxic immune cells can be used to guide the discovery of new drugs to protect the brain from deleterious immune responses.”
Investigators also noted their discoveries are relevant not only for MS, but also for other neurological, autoimmune, and infectious disorders, and that Tox-seq may be employed to discover new therapeutic candidates for different types of diseases. They also have made their toxic immune cell map publicly available for other researchers to access.
“We hope that Tox-seq will open the way to more disease-relevant transcriptomics [study of all RNA molecules found in a cell or tissue]. Information captured in genes can now be more readily related to a disease process, which will accelerate drug discovery,” Akassoglou concluded.
Leave a comment
Fill in the required fields to post. Your email address will not be published.